|
Usage Python from seqfold import fold, dg, dg_cache, dot_bracket # just returns minimum free energy dg ( "GGGAGGTCGTTACATCTGGGTAACACCGGTACTGATCCGGTGACCTCCC", temp = 37.0 ) # -13.4 # `fold` returns a list of `seqfold.Struct` from the minimum free energy structure structs = fold ( "GGGAGGTCGTTACATCTGGGTAACACCGGTACTGATCCGGTGACCTCCC" ) print ( sum ( s. Installation pyp圓 (strongly recommended) pyp圓 -m ensurepipįor a 200bp sequence (on my laptop), pyp圓 takes 2.5 seconds versus 15 seconds for CPython. Seqfold is an implementation of the Zuker, 1981 dynamic programming algorithm, the basis for UNAFold/ mfold, with energy functions from SantaLucia, 2004 (DNA) and Turner, 2009 (RNA). 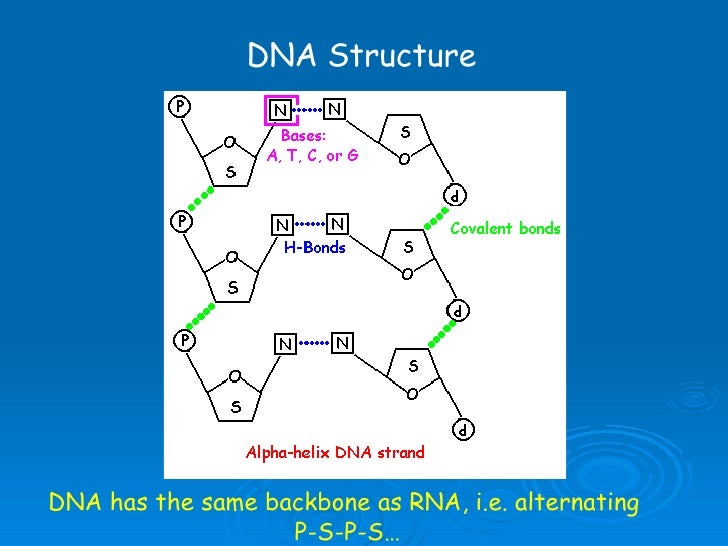
Predict the minimum free energy structure of nucleic acids.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
